library(lifelihood)
#> Loading required package: tidyverse
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ dplyr 1.2.0 ✔ readr 2.2.0
#> ✔ forcats 1.0.1 ✔ stringr 1.6.0
#> ✔ ggplot2 4.0.2 ✔ tibble 3.3.1
#> ✔ lubridate 1.9.5 ✔ tidyr 1.3.2
#> ✔ purrr 1.2.1
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ dplyr::filter() masks stats::filter()
#> ✖ dplyr::lag() masks stats::lag()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
library(tidyverse)
df <- datapierrick |>
as_tibble() |>
mutate(
par = as.factor(par),
geno = as.factor(geno),
spore = as.factor(spore),
block = rep(1:2, each = nrow(datapierrick) / 2)
)
generate_clutch_vector <- function(N) {
return(paste(
"pon",
rep(c("start", "end", "size"), N),
rep(1:N, each = 3),
sep = "_"
))
}
clutchs <- generate_clutch_vector(28)
lifelihoodData <- as_lifelihoodData(
df = df,
sex = "sex",
sex_start = "sex_start",
sex_end = "sex_end",
maturity_start = "mat_start",
maturity_end = "mat_end",
clutchs = clutchs,
block = "block",
death_start = "death_start",
death_end = "death_end",
covariates = c("par", "spore"),
model_specs = c("wei", "gam", "lgn")
)Create a lifelihoodData object
Standard errors
By default, lifelihood will not try to fit standard errors. But, you can use the se.fit argument for this purpose:
results <- lifelihood(
lifelihoodData = lifelihoodData,
path_config = use_test_config("example_config_se"),
se.fit = TRUE,
)
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_2826_9893_4331_7519/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_2826_9893_4331_7519/temp_param_range_path.txt 0 25 TRUE 0 FALSE 0 2826 9893 4331 7519 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"
summary(results)
#>
#> === LIFELIHOOD RESULTS ===
#>
#> Sample size: 550
#>
#> --- Model Fit ---
#> Log-likelihood: -343784.808
#> AIC: 687577.6
#> BIC: 687594.9
#>
#> --- Key Parameters ---
#>
#> Mortality:
#> expt_death (Intercept) -1.962 (0.072)
#> expt_death eff_expt_death_par_1 0.302 (0.078)
#> expt_death eff_expt_death_par_2 0.254 (0.084)
#> survival_param2 (Intercept) -0.210 (0.115)
#>
#> --- Convergence ---
#> All parameters within bounds
#>
#> ======================Now if we have a look at the estimations we have standard errors:
results$effects |> as_tibble()
#> # A tibble: 4 × 6
#> name estimation stderror parameter kind event
#> <chr> <dbl> <dbl> <chr> <chr> <chr>
#> 1 int_expt_death -1.96 0.0724 expt_death intercept mort…
#> 2 eff_expt_death_par_1 0.302 0.0785 expt_death coefficient_ca… mort…
#> 3 eff_expt_death_par_2 0.254 0.0841 expt_death coefficient_ca… mort…
#> 4 int_survival_param2 -0.210 0.115 survival_param2 intercept mort…Prediction
We can predict with standard errors.
- Default scale
prediction(results, "expt_death", se.fit = TRUE) |>
as_tibble() |>
sample_n(5)
#> # A tibble: 5 × 2
#> fitted se.fitted
#> <dbl> <dbl>
#> 1 -1.96 0.0724
#> 2 -1.96 0.0724
#> 3 -1.71 0.0420
#> 4 -1.96 0.0724
#> 5 -1.66 0.0293- Response scale
prediction(results, "expt_death", type = "response", se.fit = TRUE) |>
as_tibble() |>
sample_n(5)
#> # A tibble: 5 × 2
#> fitted se.fitted
#> <dbl> <dbl>
#> 1 39.9 2.53
#> 2 39.9 2.53
#> 3 39.9 2.53
#> 4 51.8 1.27
#> 5 39.9 2.53MCMC
MCMC stands for Markov Chain Monte Carlo.
Fitting with MCMC
results <- lifelihood(
lifelihoodData = lifelihoodData,
path_config = use_test_config("example_config_mcmc"),
MCMC = 30
)
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_3606_9522_1067_2829/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_3606_9522_1067_2829/temp_param_range_path.txt 30 25 FALSE 0 TRUE 0 3606 9522 1067 2829 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"
#> Warning in check_estimation(results): Estimation of 'fitness' is close to the
#> maximum bound: fitness~=999.932386277238 (bound=995). Consider increasing
#> maximum bound.Visualization
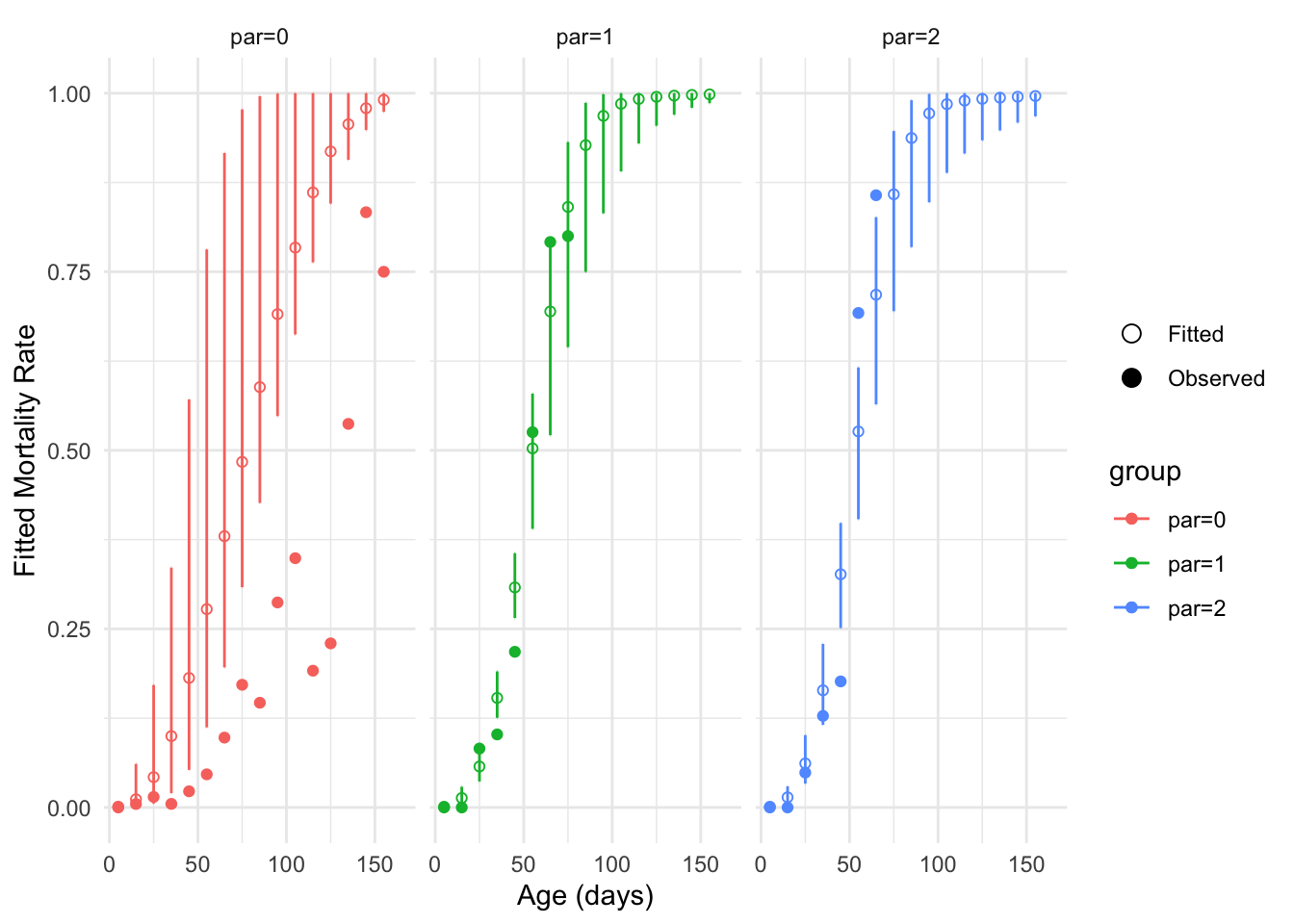
We can represent the confidence interval computed thanks to the standard errors:
plot_fitted_event_rate(
results,
interval_width = 10,
event = "mortality",
use_facet = TRUE,
groupby = "par",
xlab = "Age (days)",
ylab = "Fitted Mortality Rate",
se.fit = TRUE
)
#> Warning: Removed 20 rows containing missing values or values outside the scale range
#> (`geom_point()`).