library(lifelihood)
#> Loading required package: tidyverse
#> ── Attaching core tidyverse packages ──────────────────────── tidyverse 2.0.0 ──
#> ✔ dplyr 1.2.0 ✔ readr 2.2.0
#> ✔ forcats 1.0.1 ✔ stringr 1.6.0
#> ✔ ggplot2 4.0.2 ✔ tibble 3.3.1
#> ✔ lubridate 1.9.5 ✔ tidyr 1.3.2
#> ✔ purrr 1.2.1
#> ── Conflicts ────────────────────────────────────────── tidyverse_conflicts() ──
#> ✖ dplyr::filter() masks stats::filter()
#> ✖ dplyr::lag() masks stats::lag()
#> ℹ Use the conflicted package (<http://conflicted.r-lib.org/>) to force all conflicts to become errors
library(tidyverse)Load libraries
Fit a simple model
df <- datapierrick |>
as_tibble() |>
mutate(
par = as.factor(par),
geno = as.factor(geno),
spore = as.factor(spore)
) |>
sample_n(120)
generate_clutch_vector <- function(N) {
return(paste(
"pon",
rep(c("start", "end", "size"), N),
rep(1:N, each = 3),
sep = "_"
))
}
lifelihoodData <- as_lifelihoodData(
df = df,
sex = "sex",
sex_start = "sex_start",
sex_end = "sex_end",
maturity_start = "mat_start",
maturity_end = "mat_end",
clutchs = generate_clutch_vector(28),
death_start = "death_start",
death_end = "death_end",
covariates = c("par", "spore"),
model_specs = c("wei", "gam", "exp")
)
results <- lifelihood(
lifelihoodData,
path_config = use_test_config("config_pierrick"),
raise_estimation_warning = FALSE
)
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_3041_5709_6029_8530/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_3041_5709_6029_8530/temp_param_range_path.txt 0 25 FALSE 0 FALSE 0 3041 5709 6029 8530 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"Goodness of fit
The goodness of fit simulate datasets from a fitted model (results), refit the model on each simulated dataset (nsim), and compare simulated log-likelihood values to the original fit.
gof <- goodness_of_fit(results, nsim = 5)
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_9423_7101_7184_5681/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_9423_7101_7184_5681/temp_param_range_path.txt 0 25 FALSE 0 FALSE 0 9423 7101 7184 5681 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_8202_9851_7721_3073/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_8202_9851_7721_3073/temp_param_range_path.txt 0 25 FALSE 0 FALSE 0 8202 9851 7721 3073 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_6677_7068_362_3546/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_6677_7068_362_3546/temp_param_range_path.txt 0 25 FALSE 0 FALSE 0 6677 7068 362 3546 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_6662_2012_7512_7460/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_6662_2012_7512_7460/temp_param_range_path.txt 0 25 FALSE 0 FALSE 0 6662 2012 7512 7460 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"
#> [1] "/Users/runner/work/_temp/Library/lifelihood/bin/lifelihood-macos /Users/runner/work/Lifelihood/Lifelihood/lifelihood_4720_7660_4920_5884/temp_file_data_lifelihood.txt /Users/runner/work/Lifelihood/Lifelihood/lifelihood_4720_7660_4920_5884/temp_param_range_path.txt 0 25 FALSE 0 FALSE 0 4720 7660 4920 5884 10 20 1000 0.3 NULL 2 2 50 1 1 0.001"The goodness_of_fit() function returns an instance of class lifelihoodGOF, with the following attributes:
gof$original_loglik
#> [1] -8033.151
gof$simulated_loglik
#> [1] -11142.25 -11250.14 -10676.88 -10881.32 -10775.33
gof$n_success
#> [1] 5
gof$n_failed
#> [1] 0
gof$p_lower_or_equal
#> [1] 1You can also read the gof$fits attribute for all underlying fits. Use gof$fits[[1]] for the first one, gof$fits[[2]] for the second, and so on.
Visualization
You can use the plot() S3 method on the output of goodness_of_fit():
plot(gof)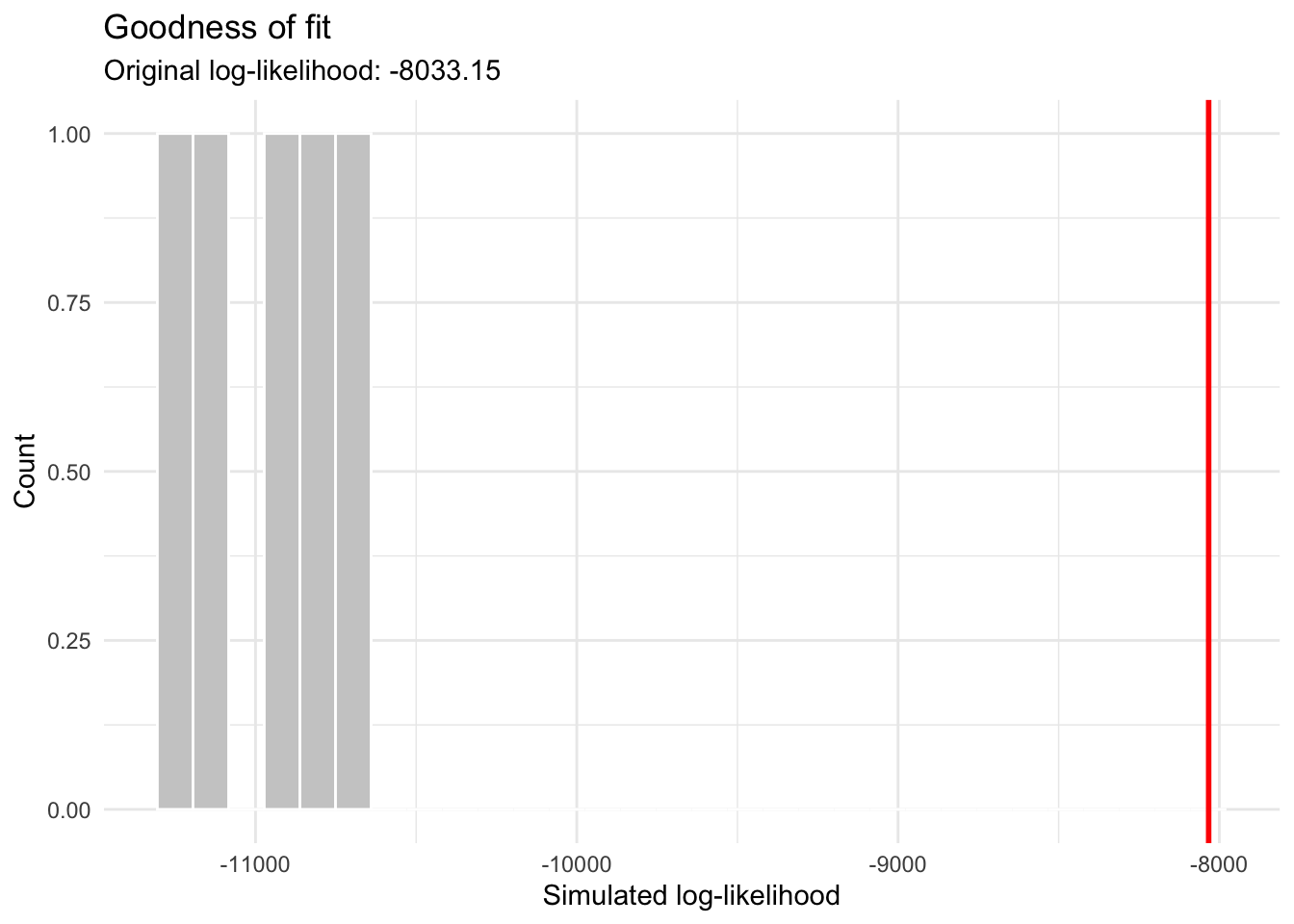
We can see here that the simulated datasets, when fitted, have a less good log-likelihood compared to the original fit. This might suggest that the original fit isn’t that great.